Research Focus
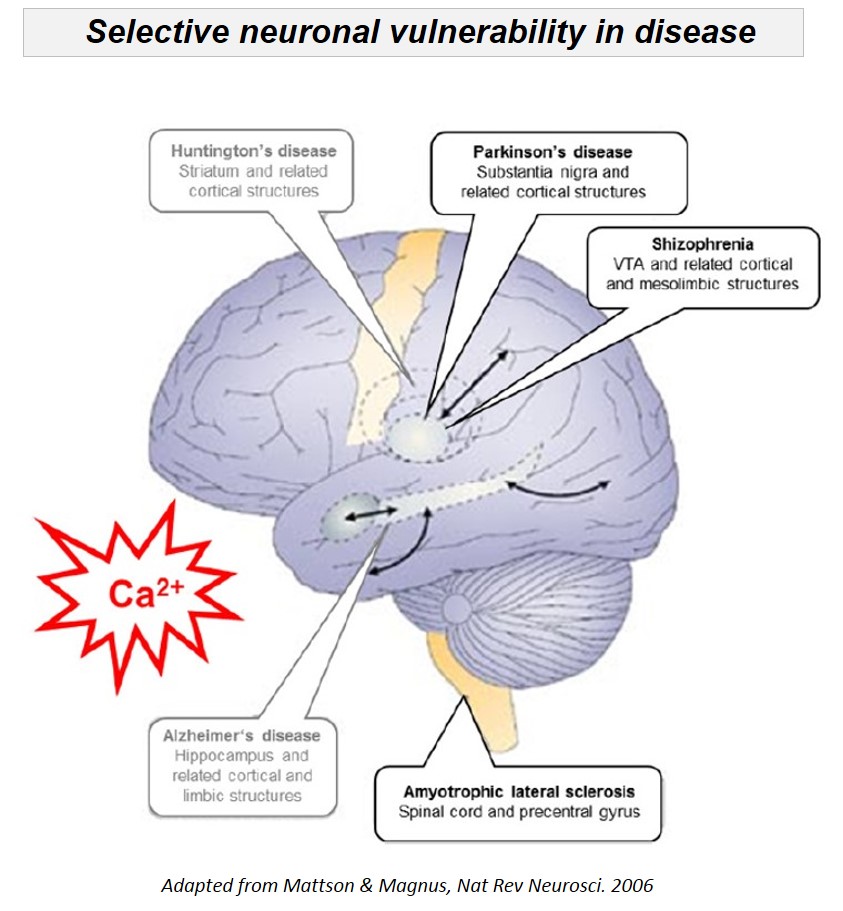
We analyze cell-specific function and gene-expression of individual neurons, to identify mechanisms of their selective (dys-)function in health and disease. Selective death or dysfunction of distinct neuronal cell populations is a main feature of many disorders of the central nervous system, like Parkinson's disease, Schizophrenia, or Amyothrophic lateral sclerosis (ALS). We focus on the dopaminergic midbrain system, as well as on analyzing cell-specific roles of ion channels, transporters and receptors (e.g. K-ATP and Ca2+ channels, dopamine and Ca2+ transporters, D2 and NMDA receptors), as their functional expression directly defines individual neuronal activity pattern and transmitter release.
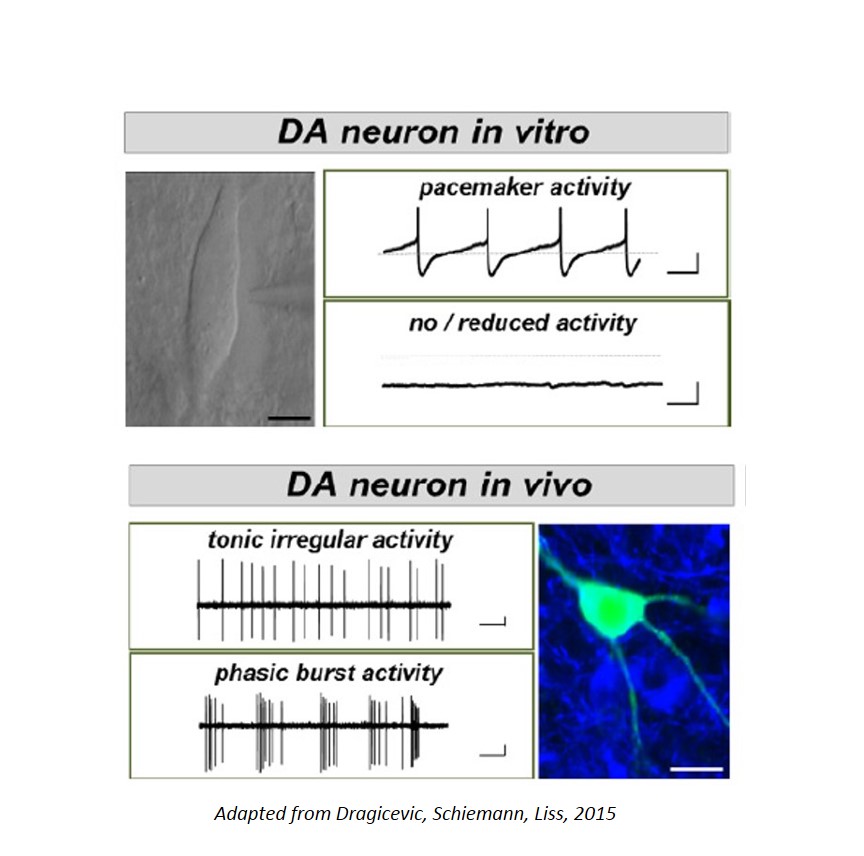
The dopamine midbrain system and the activity of dopamine releasing (DA) neurons is not only essential for motor control, but is also crucial for emotion, reward, and for cognitive functions. Moreover, subtypes of DA midbrain neurons - with different axonal projections and with distinct activity pattern - are differentially affected in diseases: e.g. Substantia nigra (SN) DA neurons are particularly prone to degeneration in Parkinson´s disease (PD), while neighboring DA neurons in the ventral tegmental area (VTA) are significantly less vulnerable to degeneration in PD, but are particularly affected in Schizophrenia and attention deficit hyperactivity disorder (ADHD). The cause for this selective vulnerability of DA neurons to distinct diseases is still unclear; however specific trigger factors, like metabolic stress, mitochondrial dysfucntion and impaired calcium homeostasis have been identified.
We aim to identify cell-specific functional and molecular differences of e.g. SN DA and VTA DA neurons, that define their distinct physiological functions, their selective transitions to disease states (e.g. malfuntion or degeneration), and their selective age-related dysfunctions.
We combine cell-specific functional analysis (e.g. retrograde tracing, in vivo & in vitro electrophysiology, MEA, calcium imaging) of wildtype and transgenic mice, as well as immunohistochemistry and laser-microdissection, with quantitative RNA and DNA analysis of individual neurons.
- in vivo tracing (using retrobeads) of mouse neurons with distinct projections.
- In vivo electrophysiology and behavioural analysis of mice.
- Brain Slice Patch-Clamp Electrophysiology and extracellular MEA recordings for functional analysis of individual neurons, as well as for cytoplasm harvest.
- UV-based Laser-Microdissektion for contact-free isolation of individual cells from tissue sections (mice and human tissues).
- Bioinformatics / database analysis for Design and analysis of gene expression experiments.
- RT-PCR und Real-time quantitative PCR (TaqMan 7900), with and without prior global Amplification for quantitaitve analysis of DNA (e.g. mitochondrial deletions), RNA and miRNA of individual cells.
- Cell-specific CGH array analysis of chromosomal DNA.
- Q-FISH analysis, Immunocytochemistry, and Stereology for cell-specific quantifcation of chromosomal telomer-length, protein expression, and for unbiased cell-number determination.
- Whole Genome Analysis for mouse and human phenotype-genotype correlations.
- MinION Access Program (MAP) for whole genome and cDNA sequencing.
Funding / grants:
- Alfried Krupp von Bohlen und Halbach-Stiftung
- BrainNet
- DFG / FWF SFB-44
- DFG Gradiertenschule CEMMA
- DFG Graduiertenschule für Molekulare Medizin
- DFG / SFB 497
- Gemeinnützige Hertiestiftung
- NGFN-plus
- The Royal Society
- Boehringer Ingelheim-Ulm University (BIU) Center
- Wellcome Trust Collaborative Award in Science
- DFG / SFB 1506
- Hamburg Institute for Advanced Study (HIAS)