High resolution transcriptomics to uncover the regulatory coordination of root stress signalling
Background: Plant roots present an almost unexplored source for enhancing plant stress resilience. Roots show a concentric organisation with epidermis, cortex, endodermis, pericycle and stele (e.g. phloem, xylem) as principal cell types. This organisation reflects partitioning of root function by cell types which provides a flexibility that secures overall root functionality under changing environments. While individual cell types differently contribute to abiotic stress signalling their function in immunity (defence against pathogens) is currently unknown. This knowledge is however essential for the generation of crops with a superior fitness under abiotic and biotic stress.
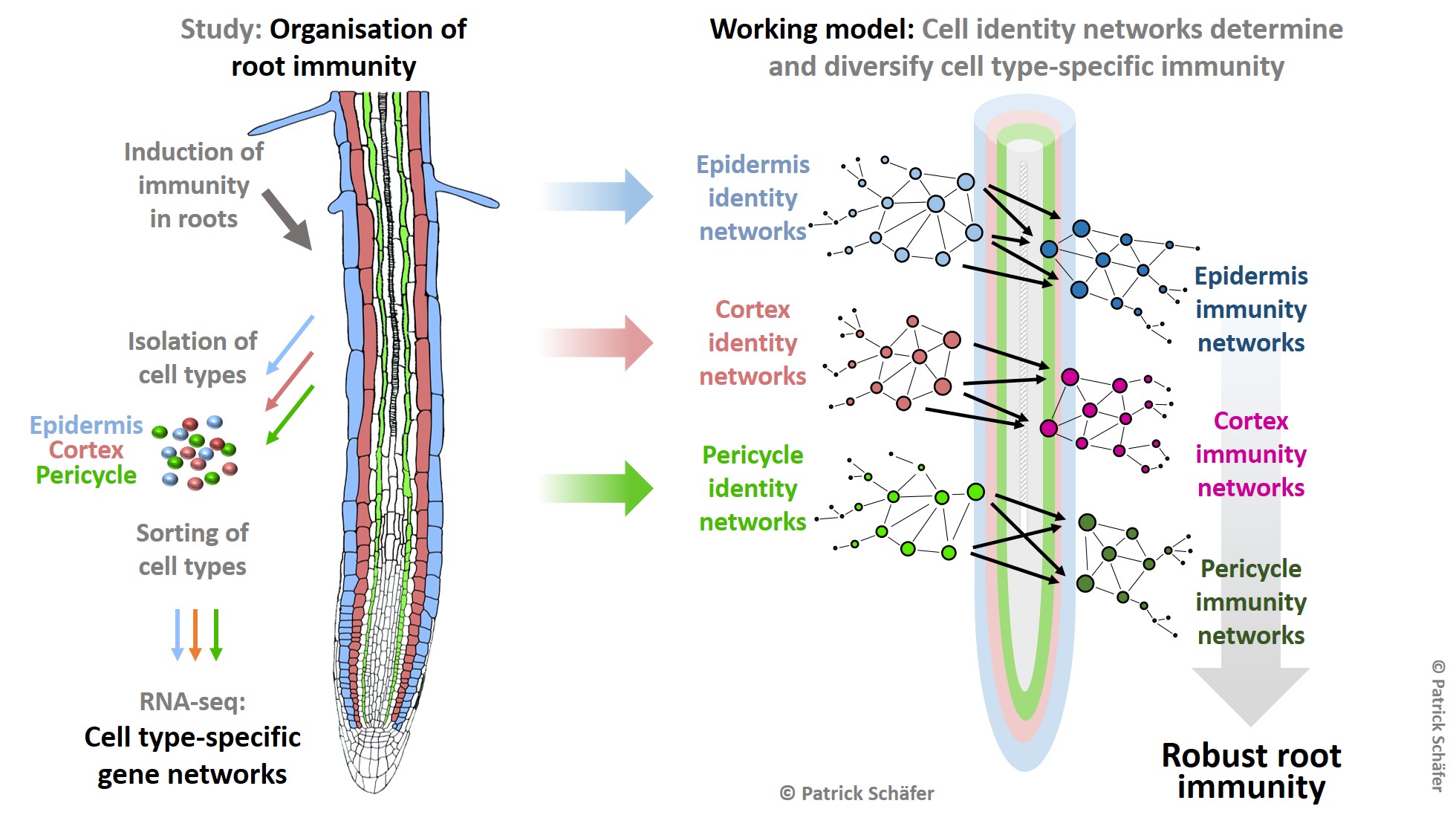
Project: To understand the organisation of immunity in root cell types, we have analysed the transcriptional networks in epidermis, cortex and pericycle cells of Arabidopsis roots treated with two known elicitors of immunity - bacterial flagellin (flg22) and the plant-derived Pep1. Our studies revealed that both elicitors activated distinct immunity networks in each cell type. To identify underlying regulatory principles, we developed a combinatorial promoter analysis tool for high accuracy prediction of functional promoter motifs. This analysis revealed pairs of transcription factors (TFs) to regulate cell type-specific immunity. Moreover, cell identity specifies cell type-specific immunity to accurately link immune response to the functional capabilities of each cell type. We want to understand how cell identity routes stress adaptation in individual cell types. Our studies revealed the significance of specific TF pair combinations in providing cell-type specifity.
Lab tools / techniques: Cell type-specific RNAseq/transcriptomics, FACS, combinatorial promoter analytics
Relevant publications:
Rich, C., Reitz, M.U., Eichmann, R., Hermann, S., Woolley-Allen, K., Esteban, E., Pasha, A., Kogel, K.H., Provart, N.J., Ott, S.1, Schäfer, P.1,2 Cell type identity determines transcriptomic immurrrne responses in Arabidopsis thaliana roots. bioRxiv doi.org/10.1101/302448.
Rich, C., Stechemesser, A.H., Finch, J., Lucas, E.S., Ott, S., Schäfer, P. (2020) Single-cell transcriptomics - a high-resolution avenue for plant functional genomics. Trends in Plant Science 25:186-197.
Lagunas, B., Achom, M., Bonyadi-Pour, R., Pardal, A.J., Richmond, B.L., Sergaki, C., Vázquez, S., Schäfer, P., Ott, S., Hammond, J.P, Gifford, M.L (2019) Regulation of resource partitioning coordinates nitrogen and rhizobia responses and autoregulation of nodulation in the legume Medicago truncatula. Molecular Plant 12: 833-846.