Research projects
RNA-Metabolism in Mitochondria of Higher Plants
Natural genetic variation affecting mitochondrial transcripts
Natural genetic variation is observed between subspecies, varieties, lines and individuals of a given species. In Arabidopsis thaliana, substantial natural genetic variation exists between different accessions (ecotypes), predominantly geographically isolated inbred populations. Natural genetic variation can affect all kinds of traits including the structure or abundance of RNA. In comparative studies of 5' and 3' extremities of major mitochondrial RNAs a numbers of 5' end polymorphisms are observed among several accessions. Their appearance in F1 reciprocals revealed that theses polymorphisms can be attributed to differences in the mitochondrial DNA or in the nuclear genome (further details see Forner et al., 2008, Stoll et al., 2013). These polymorphisms are used to identify nuclear genes required for 5' processing of mitochondrial transcripts.
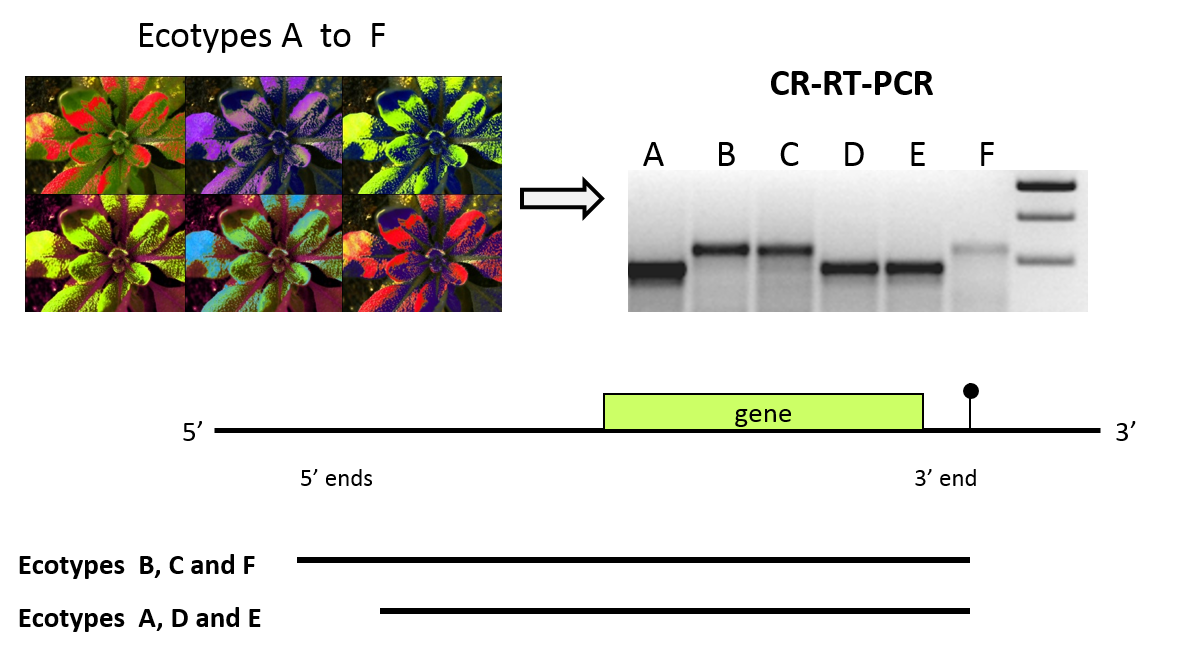
Identification and characterization of proteins required for posttranscriptional maturation of mitochondrial-encoded RNAs
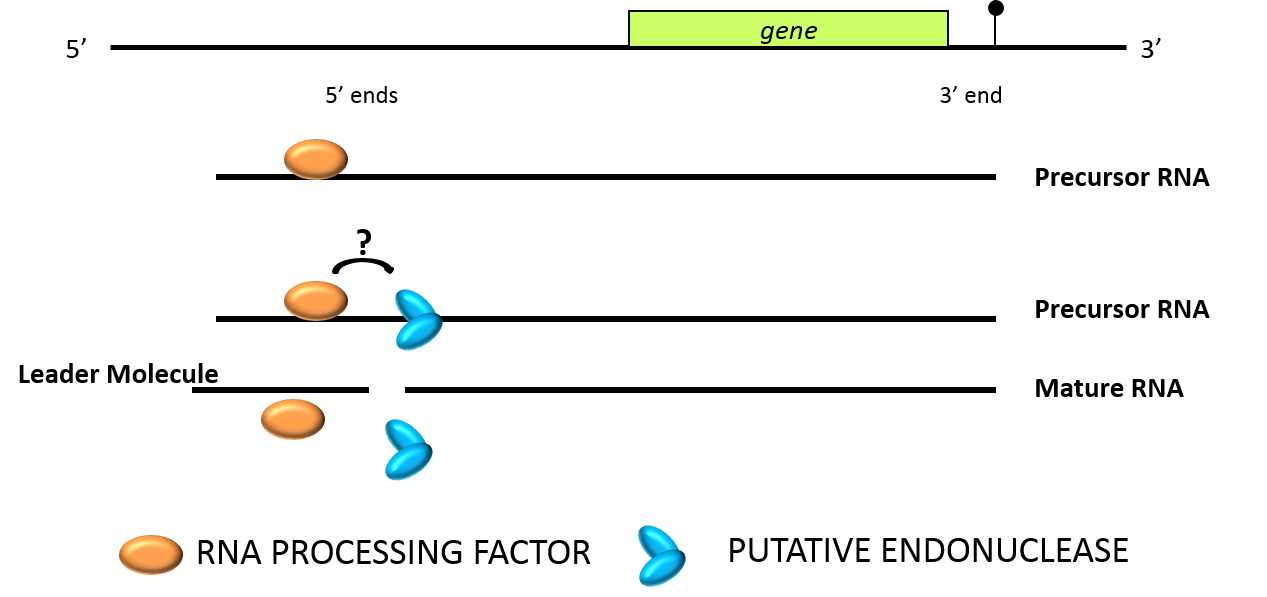
Based on natural genetic variation affecting 5' ends of mitochondrial RNAs we used a linkage analysis to identify the nuclear genes required for 5' processing of various mitochondrial transcripts in Arabidopsis thaliana (Hölzle et al., 2011; Jonietz et al., 2010; Jonietz et al., 2011; Hauler et al., 2013; Stoll et al., 2014; Stoll et al., 2015). These factors, called RNA PROCESSING FACTORs (RPF), belong to the large family of pentatricopeptide repeat proteins (PPRP) with functions in the RNA metabolism in mitochondria or (/and) chloroplasts. Some of the RPFs show high similarity to a subgroup of PPR proteins that can restore male fertility in cytoplasmic male sterile lines in other plant species. In Arabidopsis thaliana, where cytoplasmic male sterility has not been described, this class of about 27 proteins acts as specificity factors enhancing endonucleolytic generation of mature 5' ends. RPFs most likely bind specifically to the one or several target RNA directing an endonuclease to the cleavage site. Whether this requires a direct interaction between the different proteins is unclear.
Contact
Head of the research group:
Prof. Dr. Stefan Binder
Molecular Botany
Ulm University
89069 Ulm
Phone: +49 - 731-50-22625
Fax: +49 - 731-50-12-22658
stefan.binder(at)uni-ulm.de
