Publikationen 2015
Quantitative structural information from single-molecule FRET
M. Beckers, F. Drechsler, T. Eilert, J. Nagy and J. Michaelis
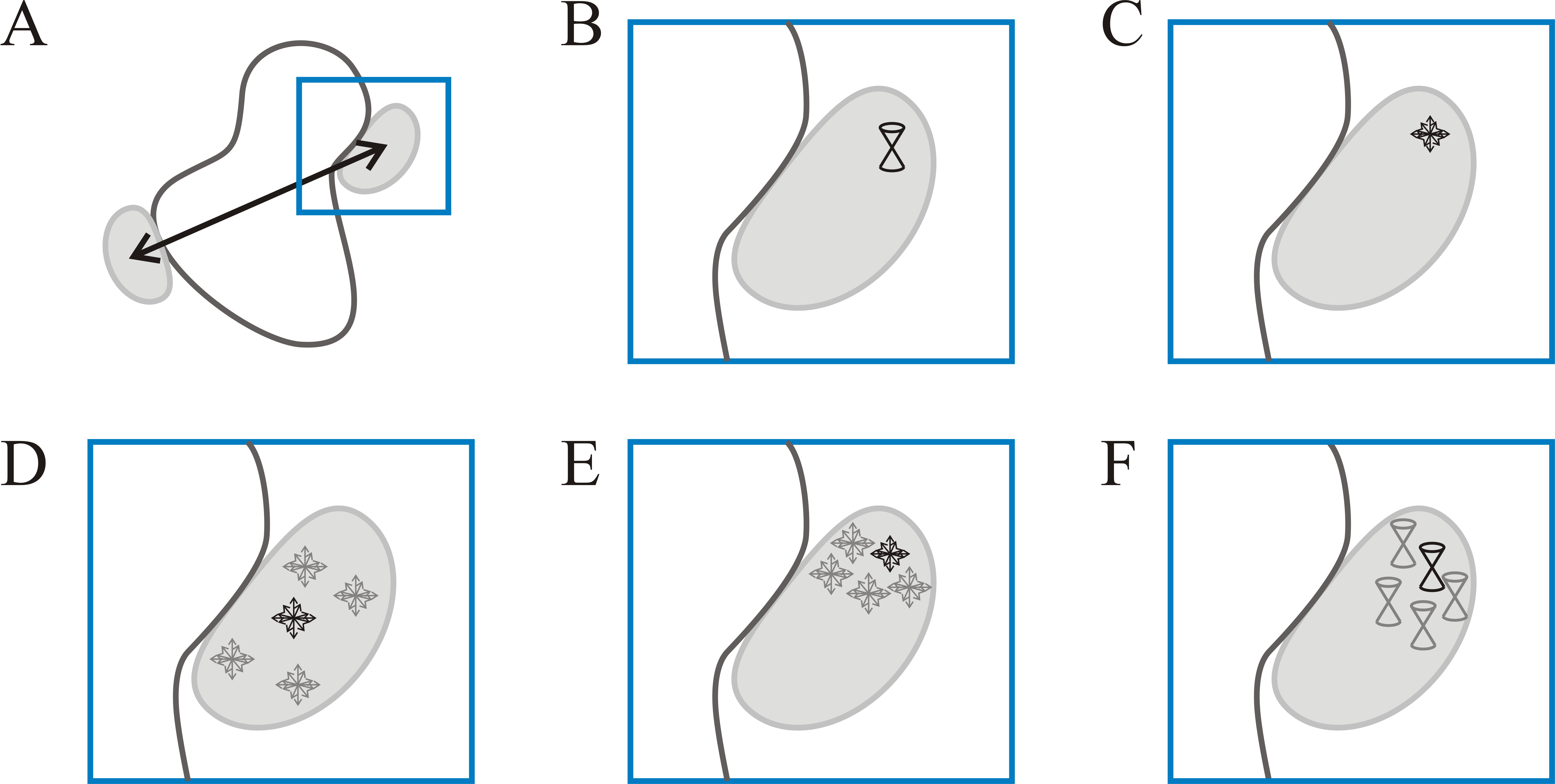
Single-molecule studies can be used to study biological processes directly and in realtime. In particular, the fluorescence energy transfer between reporter dye moleculesattached to specific sites on macromolecular complexes can be used to infer distance information. When several measurements are combined, the information can be usedto determine the position and conformation of certain domains with respect to the complex. However, data analysis schemes that include all experimental uncertaintiesare highly complex, and the outcome depends on assumptions about the state of the dye molecules. Here, we present a new analysis algorithm using Bayesian parameterestimation based on Markov Chain Monte Carlo sampling and parallel tempering termed Fast-NPS that can analyse large smFRET networks in a relatively short timeand yields the position of the dye molecules together with their respective uncertainties. Moreover, we show what effects different assumptions about the dyemolecules have on the outcome. We discuss the possibilities and pitfalls in structure determination based on smFRET using experimental data for an archaeal transcriptionpre-initiation complex, whose architecture has recently been unravelled by smFRET measurements.
Faraday Discuss., 2015, 184, 117-129
DOI: 10.1039/c5fd00110b
Complete architecture of the archaeal RNA polymerase open complex from single-molecule FRET and NPS
J. Nagy, D. Grohmann, A.C.M. Cheung, S. Schulz, K. Smollett, F. Werner, J. Michaelis
The molecular architecture of RNAP II-like transcription initiation complexes remains opaque due to its conformational flexibility and size. Here we report the three-dimensional architecture of the complete open complex (OC) composed of the promoter DNA, TATA box-binding protein (TBP), transcription factor B (TFB), transcription factor E (TFE) and the 12-subunit RNA polymerase (RNAP) from Methanocaldococcus jannaschii. By combining single-molecule Förster resonance energy transfer and the Bayesian parameter estimation-based Nano-Positioning System analysis, we model the entire archaeal OC, which elucidates the path of the non-template DNA (ntDNA) strand and interaction sites of the transcription factors with the RNAP. Compared with models of the eukaryotic OC, the TATA DNA region with TBP and TFB is positioned closer to the surface of the RNAP, likely providing the mechanism by which DNA melting can occur in a minimal factor configuration, without the dedicated translocase/helicase encoding factor TFIIH.
Nature Communications 6, Article number: 6161
DOI: 10.1038/ncomms7161
