Publikationen 2019
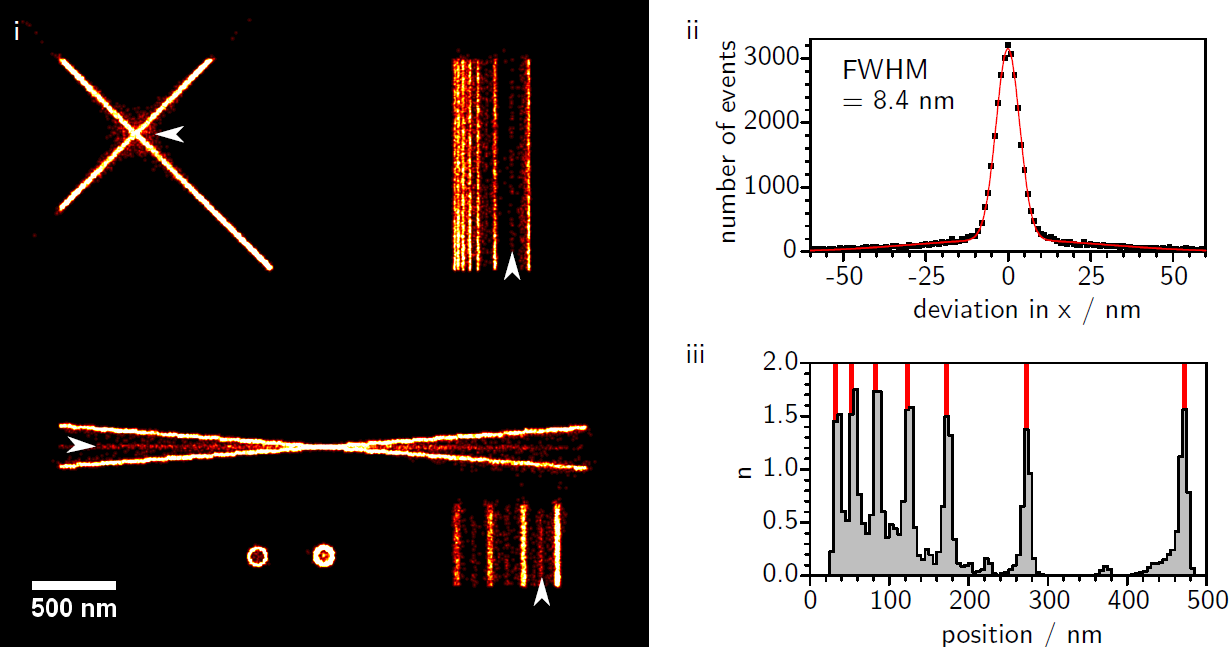
Artifact Formation in Single Molecule Localization Microscopy
Jochen M. Reichel, Thomas Vomhof, Jens Michaelis
We investigate the influence of different accuracy-detection rate trade-offs on image reconstruction in single molecule localization microscopy. Our main focus is the investigation of image artifacts experienced when using low localization accuracy, especially in the presence of sample drift and inhomogeneous background. In this context we present a newly developed SMLM software termed FIRESTORM which is optimized for high accuracy reconstruction. For our analysis we used in silico SMLM data and compared the reconstructed images to the ground truth data. We observe two discriminable reconstruction populations of which only one shows the desired localization behavior.
bioRxiv
doi.org/10.1101/700955
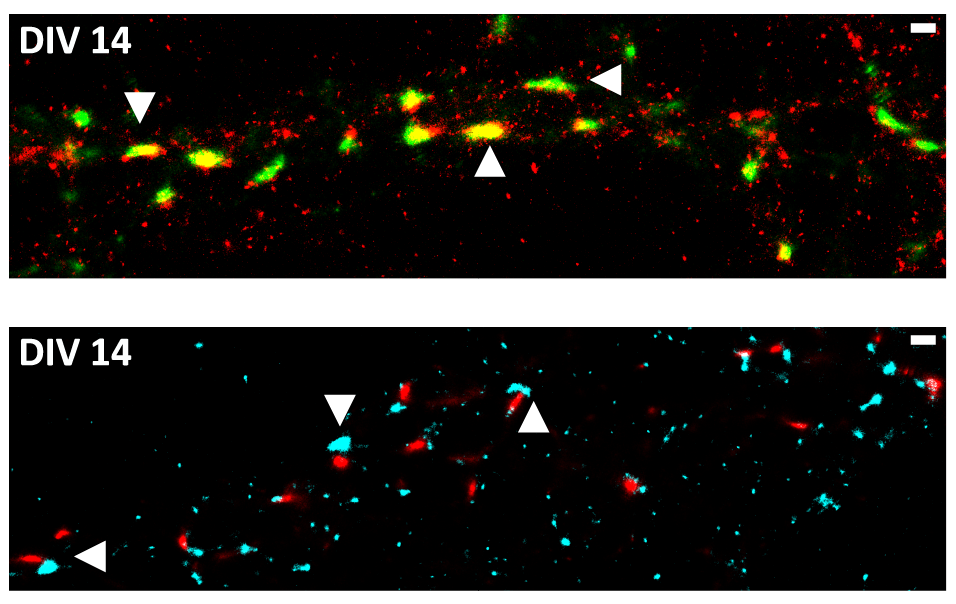
Synaptic FUS Localization During Motoneuron Development and Its Accumulation in Human ALS Synapses
Dhruva Deshpande, Julia Higelin, Michael Schoen, Thomas Vomhof, Tobias M. Boeckers, Maria Demestre and Jens Michaelis
Mutations in the fused in Sarcoma (FUS) gene induce cytoplasmic FUS aggregations, contributing to the neurodegenerative disease amyotrophic lateral sclerosis (ALS) in certain cases. While FUS is mainly a nuclear protein involved in transcriptional processes with limited cytoplasmic functions, it shows an additional somatodendritic localization in neurons. In this study we analyzed the localization of FUS in motoneuron synapses, these being the most affected neurons in ALS, using super-resolution microscopy to distinguish between the pre- and postsynaptic compartments. We report a maturation-based variation of FUS localization in rodent synapses where a predominantly postsynaptic FUS was observed in the early stages of synaptic development, while in mature synapses the protein was entirely localized in the axonal terminal. Likewise, we also show that at the synapse of human motoneurons derived from induced pluripotent stem cells of a healthy control, FUS is mainly postsynaptic in the early developmental stages. In motoneurons derived from ALS patients harboring a very aggressive juvenile FUS mutation, increased synaptic accumulation of mutated FUS was observed. Moreover increased aggregation of other synaptic proteins Bassoon and Homer1 was also detected in these abnormal synapses. Having demonstrated changes in the FUS localization during synaptogenesis, a role of synaptic FUS in both dendritic and axonal cellular compartments is probable, and we propose a gain-of-toxic function due to the synaptic aggregation of mutant FUS in ALS.
Front. Cell. Neuroscience
doi.org/10.3389/fncel.2019.00256
Mitotic chromosome binding predicts transcription factor properties in interphase
Mahé Raccaud, Elias T. Friman, Andrea B. Alber, Harsha Agarwa, Cédric Deluz, Timo Kuhn, J. Christof M. Gebhardt & David M. Suter
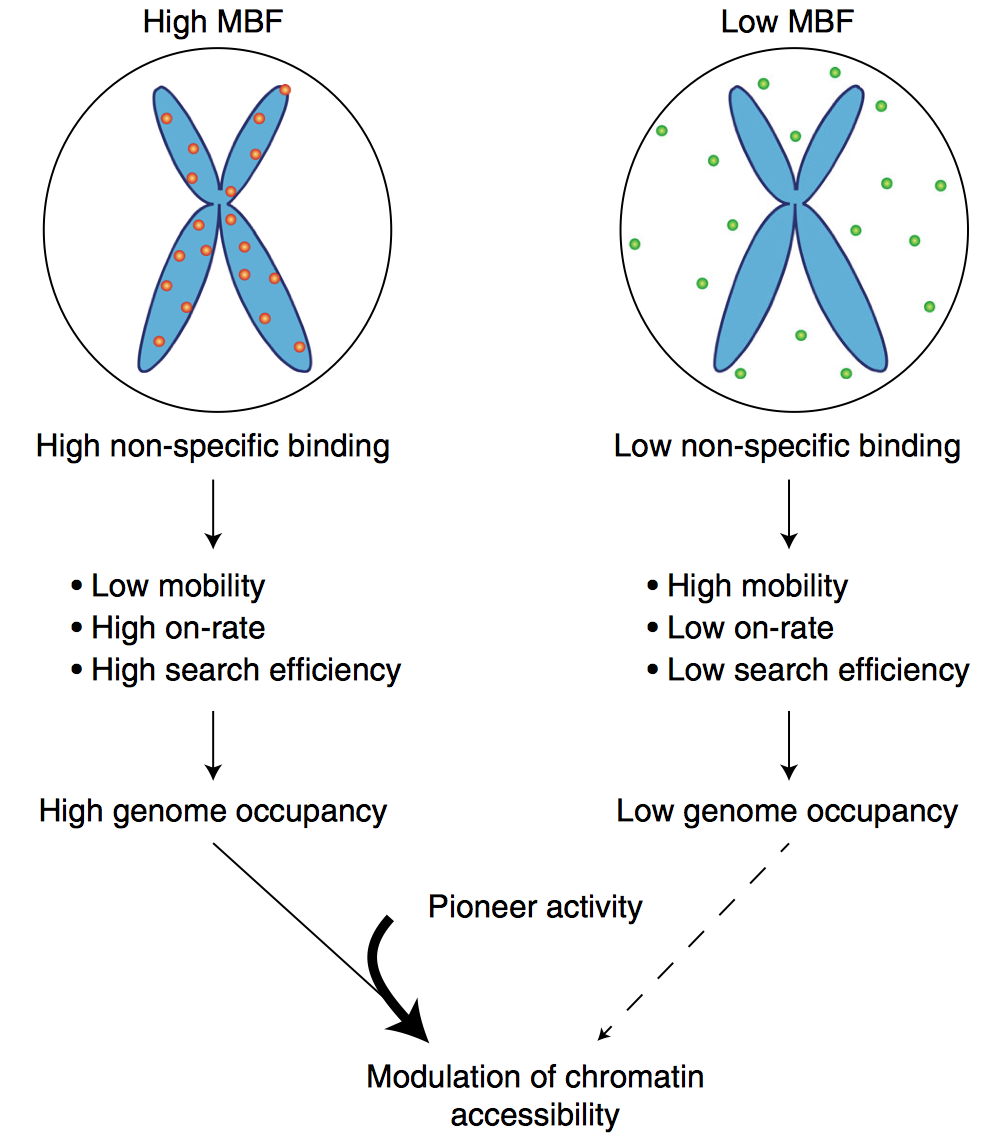
Mammalian transcription factors (TFs) differ broadly in their nuclear mobility and sequence-specific/non-specific DNA binding. How these properties affect their ability to occupy specific genomic sites and modify the epigenetic landscape is unclear. The association of TFs with mitotic chromosomes observed by fluorescence microscopy is largely mediated by non-specific DNA interactions and differs broadly between TFs. Here we combine quantitative measurements of mitotic chromosome binding (MCB) of 501 TFs, TF mobility measurements by fluorescence recovery after photobleaching, single molecule imaging of DNA binding, and mapping of TF binding and chromatin accessibility. TFs associating to mitotic chromosomes are enriched in DNA-rich compartments in interphase and display slower mobility in interphase and mitosis. Remarkably, MCB correlates with relative TF on-rates and genome-wide specific site occupancy, but not with TF residence times. This suggests that non-specific DNA binding properties of TFs regulate their search efficiency and occupancy of specific genomic sites.
Nat Commun. 2019
doi.org/10.1038/s41467-019-08417-5