Molecular mechanisms of neuronal connectivity
Projects
The differentiation of neuronal processes is essential for neuronal information processing. We are interested in proteins and microRNAs that regulate axonal branching, the elaboration of dendritic trees, and the differentiation of dendritic spines.
Several years ago we identified CALEB/NGC as a member of the EGF family of neural differentiation factors. We established a role of CALEB/NGC in regulating the complexity of dendritic trees and dendritic spines (Figure 1). Currently we are investigating the molecular mechanisms of how CALEB/NGC contributes to the differentiation of dendrites, spines, and synapses.
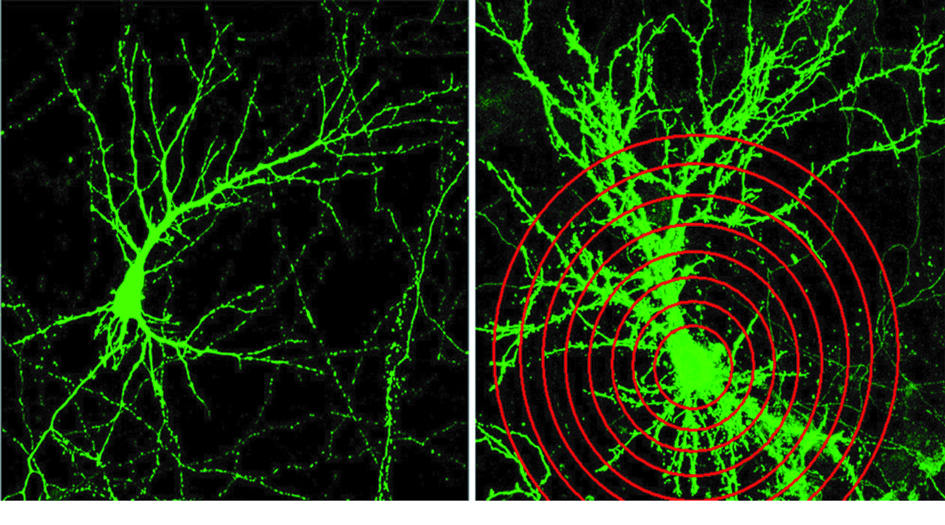
Figure 1. Primary hippocampal neurons in culture express either EGFP (left) or CALEB/NGC (right). CALEB/NGC increases the complexity of dendritic trees. Scholl analysis (red circles crossing dendrites) confirms that CALEB/NGC raises dendritic tree complexity by increasing dendritic branching.
MicroRNAs are small noncoding RNAs that regulate gene expression. They are likely to have key roles in neuronal development and plasticity. We are interested in microRNA targets that contribute to the establishment of proper neuronal connectivity. Our focus is on the function of the small GTPase RhoG, the expression of which is regulated by the microRNA miR-124. This microRNA is specifically expressed in the nervous system. RhoG and miR-124-regulated control of RhoG expression play an important role in axonal and dendritic tree differentiation.
