C03: Intrinsic heterogeneity and interactions with niches regulate trauma-induced osteoblast dedifferentiation and migration during zebrafish regeneration
PI: G. Weidinger
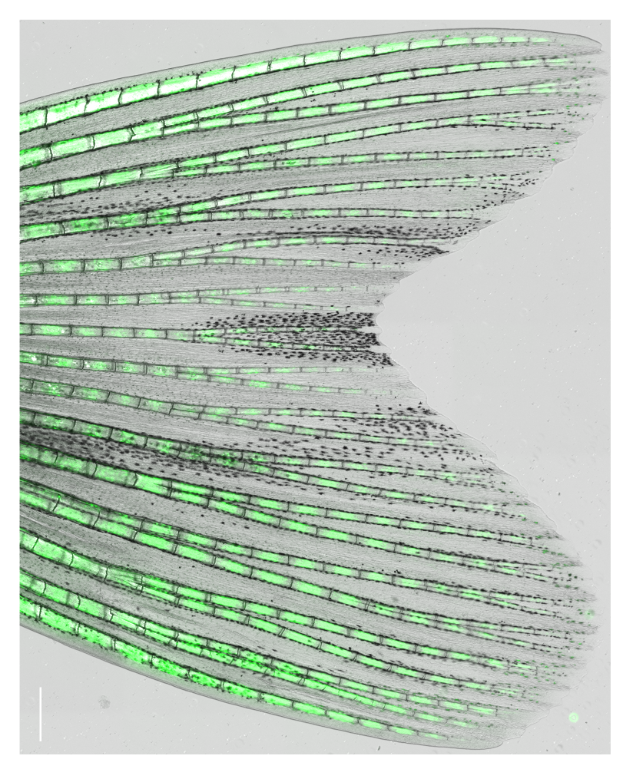
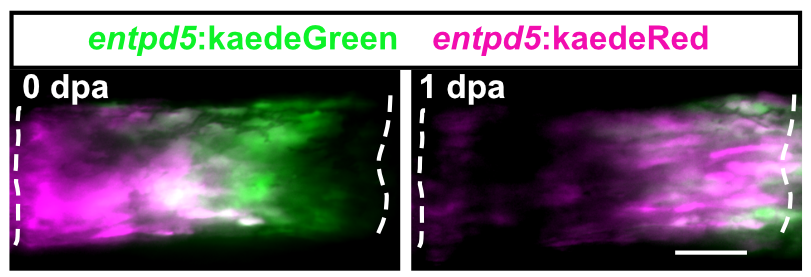
Zebrafish display highly elevated regenerative capacity compared to mammals, including an increased ability to regenerate bone. We have previously found that bone regenerates from differentiated osteoblasts which dedifferentiate to osteogenic progenitors. In the second funding period, we could show that NF-κB signaling inhibits osteoblast dedifferentiation, while EGFR signaling is required for dedifferentiation. In response to trauma, osteoblasts also become migratory. We took advantage of the ability to image osteoblast migration in live zebrafish and found that the complement system regulates osteoblast migration in vivo. Intriguingly, NF- κB and EGFR signaling do not regulate osteoblast migration, while the complement system is not required for dedifferentiation, indicating that dedifferentiation and migration are independent processes. We will further explore the interrelatedness of osteoblast trauma responses and their molecular regulation in the third funding period.
While we found that NF-κB signaling acts cell-autonomously in osteoblasts to inhibit their dedifferentiation, EGFR signaling appears to act indirectly, since the pathway is active in the basal layer of the epidermis overlying the dedifferentiating osteoblasts. This implies a novel paradigm in which interactions with other tissues regulate osteoblast responses to trauma. We plan to further explore this hypothesis in the third funding period, using genetic and transgenic methods for cell-type specific manipulation of EGFR signaling as proof-of-principle, and we will expand this idea to a potential interaction of nerves and osteoblasts, since we found that osteoblasts are also in close contact with nerve bundles.
Intriguingly, when we induce two bone defects internally in a fin, osteoblasts migrate and dedifferentiate at both injuries, while regenerative bone growth only occurs on the distal-facing side. This implies that migration and dedifferentiation represent generic osteoblast trauma responses that can occur even when they are not followed by bone regeneration. Consequently, molecular mechanisms must exist that regulate generic vs regeneration-specific responses of osteoblasts to trauma. We plan to gain molecular insight into such mechanisms in the third funding period. Finally, we have discovered that osteoblasts appear to be heterogenous during bone homeostasis and in the extent by which they dedifferentiate in response to trauma. This could be an indication of the existence of unrecognized osteoblast subpopulations. We will resolve osteoblast heterogeneity and molecular transition stages during dedifferentiation using single cell RNA sequencing.
Projektleiter
Prof. Dr. Gilbert Weidinger
Institute of Biochemistry and Molecular Biology
Ulm University
M24, Level 3, Room 323
Albert-Einstein-Allee 11
89081 Ulm
Tel.: +49 731 500 23290
Fax: +49 731 500 23277
gilbert.weidinger(at)uni-ulm.de
www.uni-ulm.de/weidinger